The Ulrich and the Beli lab collaborate to describe an exciting new tool to reprogramme linkage topologies of polyubiquitin chains
07-03-2022
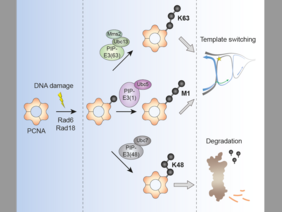
The posttranslational protein modifier ubiquitin is involved in most aspects of the cellular metabolism. Its ability to form polymeric chains of varying topologies contributes to its versatility as a signaling molecule. However, the significances of the different linkage topologies are poorly understood due to the lack of suited tools to study them. The Ulrich lab now describes an enzyme-based strategy of ubiquitin linkage reprogramming in vivo, designed to explore the functional relevance of polyubiquitin chain linkages in a defined biological context. The researchers describe that reprogramming of polyubiquitilyation of the budding yeast replication factor PCNA in response to DNA damage alters the fate of the modified protein and therefore the cellular consequences. Proof-of-principle experiments show that the method can be extrapolated to other proteins, which opens up new exiting possibilities to explore the function of polyubiquitin chains on a myriad of other ubiquitin-modified proteins. To read the full paper click here.
Wegmann S, Meister C, Renz C, Yakoub G, Wollscheid HP, Takahashi DT, Mikicic I, Beli P, Ulrich HD (2022) Linkage reprogramming by tailor-made E3s reveals polyubiquitin chain requirements in DNA-damage bypass. Mol Cell 82(8):1589-1602, doi: 10.1016/j.molcel.2022.02.016 Link